Universal Multiplexable matK Primers for DNA Barcoding of Angiosperms
- Background and Aims: PCR amplification of the matK barcoding region is often difficult when dealing with multiple angiosperm families. We developed a primer cocktail to amplify this region efficiently across angiosperm diversity.
-
Methods and Key Results: We developed 14 matK primers (seven forward, seven reverse) for multiplex PCR, using sequences available in GenBank for 178 taxa belonging to 123 genera in 41 families and 18 orders. Universality of these new multiplexed primers was tested with 53 specimens from 44 representative angiosperm families in 23 different orders. Our primers showed high PCR amplification and sequencing success.
-
Conclusions: These results show that our newly developed primers are highly effective for multiplex PCR and can be employed in future barcode projects involving taxonomically diverse samples across angiosperms. Using multiplex primers for barcoding will reduce the cost and time needed for PCR amplification.
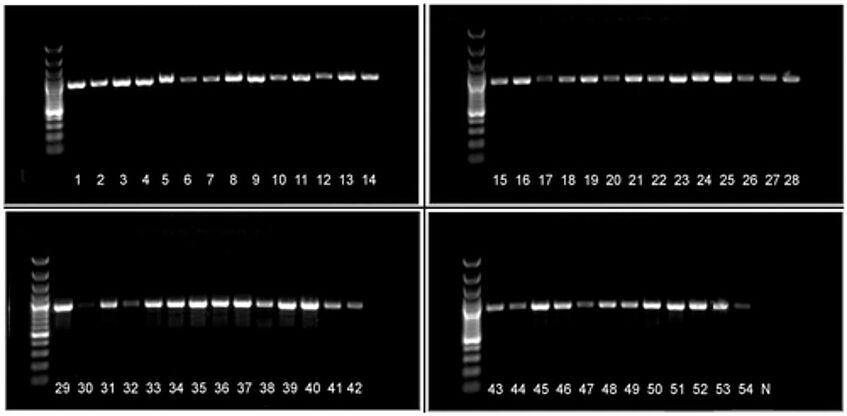
Images of PCR amplicons for representatives of 53 angiosperm families using multiplex PCR with the newly developed degenerate primers (matK-413f-1 to matK-413f-5, matK-1227r-1 to matK-1227r-5). Bands are approximately 900 bp. Most of the samples were amplified using 2× ReddyMix. Low-quality DNA samples (slot 30) that failed PCR could be amplified using 2× Phusion Green HS II Hi-Fi PCR Master Mix (slot 31). For detailed sample description, see Table 3. Ladder: GeneRuler 100 bp Plus DNA Ladder (#SM0321; Thermo Fisher Scientific, Waltham, Massachusetts, USA). N = negative control.
